|
Biology of the Malaria Parasite, Plasmodium
Almost half of the world's population is at risk of malaria (https://www.who.int/features/factfiles/malaria/en/). It kills over 400,000 people each year and infects more than 200 million more. In addition, malaria imposes a significant economic cost on endemic countries contributing to their poverty.
Malaria infection begins with the transmission of Plasmodium sporozoites from an infected mosquito to humans (Figure 1). Sporozoites invade hepatocytes in the host liver by forming a vacuole within which they develop further into 'liver stages'. Infection by sporozoites of the liver is the first key step in establishing a robust infection in the host that ensures its subsequent propagation in the blood stream. My research focuses on identifying key Plasmodium molecules required for the parasite's infection of the liver. We utilize a combination of molecular genetics, cell biology, imaging and genomics to identify parasite proteins required for successful infection of, development within and exit from mammalian hepatocytes.
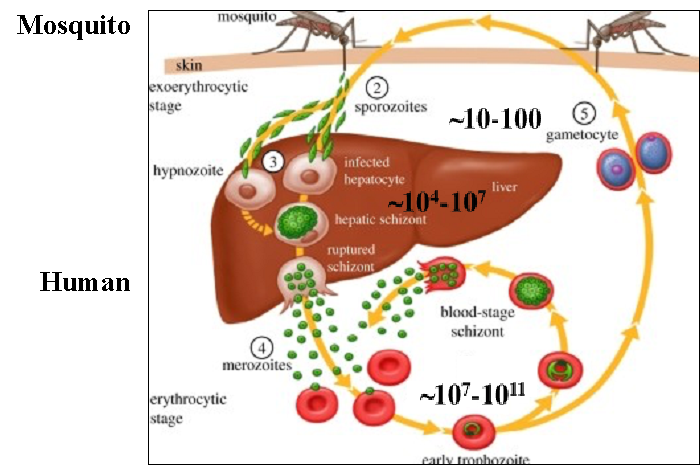
Figure 1: Plasmodium infection (adapted from Hill 2011, Philos Trans R Soc Lond B Biol Sci) begins with the transmission of parasite forms called 'sporozoites'to the mammalian host through mosquito bites. Sporozoites infect hepatocytes wherein they undergo a dramatic increase in parasite numbers - from only ~10-100 transmitted by the mosquito to ~104-107 that exit the liver. Parasites released from the liver (liver stages) initiate repeated rounds of invasion of erythrocytes and commence the pathology-inducing step of malaria infection. Preventing the parasite's infection of the liver can attenuate or stop malaria infection at an early step of the cycle.
A comprehensive understanding of the biological mechanisms that govern sporozoite infection and liver stage development (together known as the pre-erythrocytic cycle) will enable better chemical prophylaxis strategies against malaria. Towards this end, my research has focused on identifying parasite kinases that play crucial roles in liver infection and could be potentially targeted by anti-parasitic drugs. My research program aims to (1) understand the function of Plasmodium second messenger signaling pathways during the first key step of a mammalian infection, (2) validate select parasite kinases as potential drug targets and (3) develop early hit-to-lead programs for inhibitors of validated kinases.
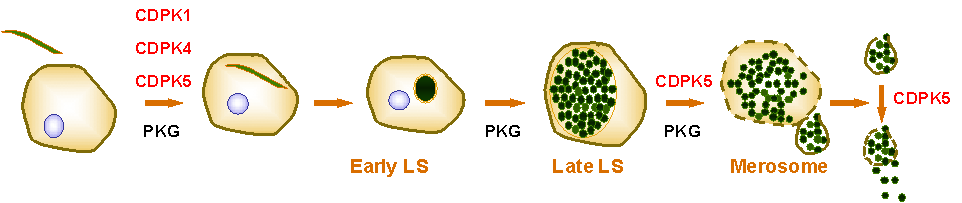
Figure 2: Plasmodium kinases that play a critical role during the parasite's infection of the liver. These include PKG and members of the CDPK family. Sporozoites invade hepatocytes and develop into liver stages (LS). LS exit the hepatocyte in membrane-bound vesicles termed 'merosomes'. Merosomes are deposited into the bloodstream, where they disintgrate and release LS. Released LS then infect erythrocytes. Our work has demonstrated that PKG is required for the parasite's invasion of hepatocytes and the development of merosomes. In addition, it has established that CDPK1, 4 and 5 have partially overlapping but non-redundant roles during parasite entry, and that CDPK5 is required for merosome formation and release of LS from merosomes.
PLASMODIUM PKG IS ESSENTIAL FOR INFECTION. A primary focus on my research has been the parasite's cGMP dependent protein kinase (PKG). These studies have established that (1) cGMP-dependent signaling is crucial in parasite development (2) parasite PKG is essential for parasite entry and exit from the host liver (3) parasite PKG function during the parasite's liver cycle is chemically "targetable". funded work on PKG, we are extending our studies by defining the mechanism of Plasmodium PKG action. We have identified potential direct substrates of the kinase by defining its unique consensus phosphorylation site and identifying parasite proteins that physically interact with it, in vitro. This work strongly suggests that Plasmodium PKG phosphorylates a subunit of the 19S regulatory particle of the parasite's 26S proteasome. It points to an intriguing intersection between PKG and the parasite's proteasomal machinery.
A TRIO OF CALCIUM DEPENDENT PROTEIN KINASES (CDPKS) IS REQUIRED FOR SPOROZOITE MOTILITY. We have studied a family of Ca+2 dependent protein kinases (CDPKs) present in Plasmodium. Ca+2 signaling is important for both parasite entry into and egress from hepatocytes. An indication of the variety of physiological roles played by Ca+2 comes from the large number of CDPK homologs present in Plasmodium. Using a combination of chemical biology, classical genetics and biochemical tools, we are dissecting the overlapping functions of different family members in in sporozoite and liver stage biology. We have demonstrated that CDPK1, 4 and 5 play an important role in sporozoite motility and entry into hepatocytes. A small molecule inhibitor of CDPK4 blocks sporozoite motility and invasion into hepatocytes. Liver stage parasites treated with this CDPK4 inhibitor or missing the CDPK4 gene are partially inhibited in their exit from the liver. We have identified that CDPK4 is complemented by at least two additional family members, CDPK1 and CDPK5. Loss of either CDPK1 or CDPK5 decreases sporozoite motility and invasion, much like the loss of CDPK4. In addition, CDPK5 is required for parasite exit. One or more of these kinases may be validated as targets for drug discovery.
|
||||||


 Purnima Bhanot, Ph.D.
Purnima Bhanot, Ph.D.