Email: mandecwl@njms.rutgers.edu |
Biomolecular Interactions in Ribosomal Translation
Search for antibiotic drug leads that inhibit protein synthesis through studying interactions between EF-Tu and tRNA
One initial crucial step of protein biosynthesis in all known organisms is the assembly of a ternary complex consisting of elongation factor Tu (EF-Tu), an aminoacylated transfer RNA molecule (aa-tRNA), and guanosine triphosphate (GTP). The ternary complex is the vehicle for delivery of amino acids to the growing protein chain, bringing the aa-tRNA into the codon-programmed A-site of the ribosome. About half of all antibiotics target bacterial protein synthesis, but there are no inhibitors currently in use in clinical practice that target this particular step. A few known inhibitors that are active in vitro (kirromycin and others) are not effective against gram-negative bacteria in a clinical setting because they do not enter bacterial cells.
By modifying E. coli EF-Tu genetically, and with attachment of fluorescent dyes to the EF-Tu and an E. coli tRNA molecule specific for the amino acid phenylalanine (tRNAPhe), we have been able to demonstrate fluorescence resonance energy transfer (FRET) upon assembly of the ternary complex. Further, the modified molecules appear to be capable of carrying out their function in protein biosynthesis normally in vitro. These reagents are thus a powerful tool for rapid high throughput screening (HTS) of a library of small molecules in search for inhibitors of ternary complex formation. The inhibitor will reduce fluorescence transfer observed in the assay. Our work is aimed at identification of drug leads for antibiotics that target protein synthesis at a different step than all other antibiotics currently in use.
The program has roots in the work done in our laboratory under an NIH program on "Revolutionary Genome Sequencing Technologies - the $1000 Genome". Under this grant, we established the principles of the FRET assay involving EF-Tu and tRNA.
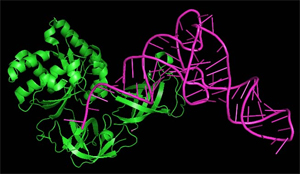
EF-Tu (green) interacts with tRNA (pink) on the ribosome (not shown). Data from 1mj1.pdb file.
Other Research interests
In addition, Dr. Mandecki's research interests include the mechanisms of frameshifting in ribosomal translation, phage display, protein structure and function, and innovative techniques in nucleic acid and protein analysis, including assay development for diagnostics in vitro. Dr. Mandecki is recognized for the assembly in the laboratory of the longest (at the time) synthetic DNA molecule (plasmid pWM521) (1990). Dr. Mandecki has made many contributions to biotechnology and biophysics. He is the inventor of a small electronic chip ("p-Chip") capable of wireless communication currently used to tag small laboratory animals. The p-Chip has also been implanted in an embryo.
Collaborators
The project is a collaboration with Dr. Emanuel Goldman at the Department. We also interact with many colleagues at the Department and NJMS: Drs. Hieronim Jakubowski, Hong Li, Sanjay Tyagi and many others.
Dr. Mark Henderson, NCATS-NIH. Arrangements have been made to collaborate and to perform an HTS aimed at identification of inhibitors of the EF-Tu:tRNA interaction at NCATS.
Biosketch
Dr. Wlodek Mandecki received a B.S. in physics from the University of Warsaw, Poland, and a Ph.D. in biochemistry from the Institute of Biochemistry and Biophysics of the Polish Academy of Sciences. Dr. Mandecki performed his postdoctoral training at the University of California at Los Angeles, the University of Wisconsin at Madison, and the University of Colorado at Boulder. He worked at Abbott Laboratories, a pharmaceutical company in Chicago, IL, for 12 years as an R&D Manager. His appointments in the biotechnology field include working as Director of Molecular Biology at DGI Biotechnologies (Edison, NJ). He is the Founder of a successful start-up company, PharmaSeq, Inc. (located near Princeton, NJ), recently acquired by p-Chip Corporation.
Dr. Mandecki has made many important contributions in the area of gene regulation, expression and mutagenesis, protein structure-function, protein design and engineering, phage display and drug discovery. He invented a light-activated electronic microtransponder ("p-Chip"), used for tagging of laboratory animals. Several biotechnology applications are being developed. Dr. Mandecki’s work has been funded by grants from the U.S. National Institutes of Health and the National Science Foundation. Dr. Mandecki has more than 70 scientific papers in genetics, molecular biology and diagnostics, and over 30 patents.
Published Work in MyBibliography (NCBI/NIH)
Selected Publications
-
Løset GÅ, Mandecki W, Sandlie I (2021) Phage display and selection of protein ligands. In Practical Handbook of Microbiology, 4th Edition (eds. L. Green, E. Goldman), CRC Press, Boca Raton, FL, chapter 9
-
Bhatt R, Chudaev M, Mandecki W, Goldman E. Engineered EF-Tu and tRNA-based FRET screening assay to find inhibitors of protein synthesis in bacteria (2018) Assay Drug Dev Technol. May/Jun;16(4):212-221
-
Shah S, Mandecki W, Li J, Gryczynski Z, Borejdo J, Gryczynski I, Fudala R (2016) FRET study in oligopeptide-linked donor–acceptor system in PVA matrix. Methods Appl Fluoresc 4:047002
-
Mandecki W, Rodriguez EF, Drawbridge J (2016) Tagging of individual embryos with electronic p-Chips. Biomed Microdevices 18(6):100
-
Li J, Veltri RW, Yuan Z, Christudass CS, Mandecki W (2015) Macrophage inhibitory cytokine 1 biomarker serum immunoassay in combination with PSA is a more specific diagnostic tool for detection of prostate cancer. PLoS ONE 10(4):e0122249
-
Liu W, Kavaliauskas D, Schrader JM, Poruri K, Birkedal V, Goldman E, Jakubowski H, Mandecki W, Uhlenbeck OC, Knudsen CR, Goldman YE, Cooperman BS (2014) Labeled EF-Tus for rapid kinetic studies of pretranslocation complex formation. ACS Chem Biol. 9(10):2421-243
-
Chudaev M, Poruri K, Goldman E, Jakubowski H, Jain MR, Chen W, Li H, Tyagi S and Mandecki W (2013) Design and properties of efficient tRNA:EF-Tu FRET system for studies of ribosomal translation. Protein Eng Des Sel 26:347-357
-
Bharill S, Chen C, Stevens B, Kaur J, Smilansky Z, Mandecki W, Gryczynski I, Gryczynski Z, Cooperman BS, Goldman YE (2011) Enhancement of single-molecule fluorescence signals by colloidal silver nanoparticles in studies of protein translation. ACS Nano 2011 5:399-407
-
Song, L., Mandecki, W., Goldman, E. (2003) Expression in E. coli of non-ORF sequences isolated from phage display: upstream start and stop required for translation reinitiation. FASEB J. 17: 1674-1681
-
Zemsky, J., Mandecki W., and Goldman, E. (2002) Genetic analysis of translation in the -1 frame of an unusual non-ORF sequence isolated from phage display. Gene Expression 10: 109-114
-
Goldman, E., Korus, M. and Mandecki, W. (2000) Efficiencies of translation in three reading frames of unusual non ORF sequences isolated from phage display. FASEB J. 14: 603-611
-
Carcamo, J., Ravera, M.W., Brissette, R., Dedova, O., Beasley, J., Alam-Moghi, A., Wan, J., Blume, A. and Mandecki, W. (1998) Unexpected frameshifts from gene to expressed protein in a phage-displayed peptide library. Proc. Natl. Acad. Sci. USA. 95: 11146-11151
-
Brennan, C.A., Christianson, K., LaFleur, M.A. and Mandecki, W. (1995) A molecular sensor system based on genetically engineered alkaline phosphatase. Proc. Natl. Acad. Sci. U.S.A. 92: 5783-5787
-
Brennan, C., Christianson, K., Surowy, T. and Mandecki, W. (1994) Modulation of enzyme activity by antibody binding to an alkaline phosphatase-epitope hybrid protein. Protein Engineering 7: 509-514
-
Mandecki, W., Shallcross, M.A., Sowadski, J. and Tomazic-Allen, S. (1991) Mutagenesis of conserved residues within the active site of Escherichia coli alkaline phosphatase yields enzymes with increased kcat. Protein Engineering 4: 801-804
-
Mandecki, W., Hayden, M., Shallcross, M.A. and Stotland, E. (1990) A totally synthetic plasmid for general cloning, expression and mutagenesis in Escherichia coli. Gene 94: 103-107


 Wlodek Mandecki, Ph.D.
Wlodek Mandecki, Ph.D.